
This is a relatively new format that is very similar to BAM as it also retains the same information as SAM and is compressed, but it is much smarter in the way that it stores the information. Some special tools are needed in order to make sense of BAM, such as Samtools, Picard Tools, and IGV which will be discussed in some of the latter sections. This is the same format except that it encoded in binary which means that it is significantly smaller than the SAM files and significantly faster to read, though it is not human legible and needs to be converted to another format (i.e. HWI-ST865:416:C6CG0ACXX:1:1113:14118:89232 16 I 15 1 100M * 0 0 CTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAA AS:i:0 XS:i:0 XN:i:0 XM:i:0 XO:i:0 XG:i:0 NM:i:0 MD:Z:100 YT:Z:UU HWI-ST865:416:C6CG0ACXX:1:1215:16359:6484 16 I 9 1 85M * 0 0 CTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCC EEEEFFFFDAGHHHIJJIIJJJJJIJJJJIJIJJIGIIJJJJJJJIJJJJJJJIIJJJIIJGJJIJJJJJJJHFHHHFFFFFCCB AS:i:0 XS:i:0 XN:i:0 XM:i:0 XO:i:0 XG:i:0 NM:i:0 MD:Z:85 YT:Z:UU

HWI-ST865:416:C6CG0ACXX:1:1313:9073:43827 0 I 2 1 99M * 0 0 CCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCTAAGCCT AS:i:0 XS:i:0 XN:i:0 XM:i:0 XO:i:0 XG:i:0 NM:i:0 MD:Z:99 YT:Z:UU The definition for the operators can be found here: Example SAM file Instead of writing the whole alignment out, operators have been defined and are used in combination with numbers to explain which part of the sequence aligns, which doesn’t, and everything in between. This is a shorthand way to encode an entire alignment. Different algorithms report it differently but nonetheless, the greater the number the better the alignment (generally). This value reports how well the read aligned to the reference. To make it easy you can check here to either encode or decode a bitwise flag. One important thing to note is that any combination of these flags results in one integer, which makes interpreting it a bit difficult. and any combination of the available tags, seen below: It tells you whether the read aligned, is marked a PCR duplicate, if it’s mate aligned, etc.
#Bam file format nh tag code#
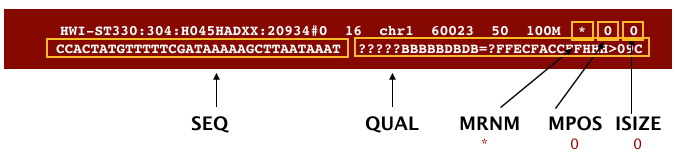
The bitwise flag is a lookup code to explain certain features about the particular read (exact same concept as Linux permission codes!). Let’s look at some of the fields that aren’t very self explanatory: Bitwise Flag The descriptions for them can be found below: Field DescriptionsĮach row contains 11 mandatory fields.
#Bam file format nh tag full#
The full list of available header fields can be found below. Some example information that can be entered into the header is: command that generated the SAM file, SAM format version, sequencer name and version. The header varies in size but adheres to a particular format depending on what information you decide to add. It consists of a header, a row for every read in your dataset, and 11 tab-delimited fields describing that read. This is generated by almost every alignment algorithm that exists. This is the most basic, human readable format of the three. With this format not only is the alignment retained but the associated quality scores (both mapping and base quality), the original read itself, paired-end information, sample information, and many more features. The first of these to be introduced was Sequence Alignment Map (SAM). Initially there were many different formats, most of them proprietary, which were space inefficient and either held too much or too little information. These formats were introduced to standardize how alignments are reported. The official SAM documentation can be found here.

#Bam file format nh tag install#
Instructions to install R Modules on Dalma.Salmon & kallisto: Rapid Transcript Quantification for RNA-Seq Data.Over-Representation Analysis with ClusterProfiler.Gene Set Enrichment Analysis with ClusterProfiler.NGS Sequencing Technology and File Formats.Next-Generation Sequencing Analysis Resources.
